Workshop
Hobart, Australia 1-5 April 2019
Organiser: Oliver Berry, CSIRO Environomics Future Science Platform.
The increasingly easy access to large SNP datasets for wild organisms means that it’s an exciting time to be working in population genomics and molecular ecology. The size of these datasets opens up a wealth of possibilities for deeper and more diverse analyses and understandings of biological process.
This workshop covered a range of topics to do with analysing SNP data generated by representational sequencing approaches like RAD, ddRAD, DArTSeq and others. The workshop began with an introduction to getting one's data into standard format for pre-processing using packages like Radiator (Thierry Gosselin, Montreal) and dartR (Arthur Georges, UCanberra). Lots of hands on with real datasets, and discussion of the nuances of filtering for purpose.

Participants were introduced the options for filtering data on such things as Call Rate, Repeatability, taking out secondaries (multiple SNPs per sequencing tag) etc. Peter Grewe (CSIRO) gave a presentation on the importance of quality control when collecting samples.
The second day focussed on clustering strategies, led by Scott Foster (CSIRO), and using stockR for examining structure within a SNP dataset. Jeff Good (U Montana) gave a seminar on using natural history collections to better understand adaptive responses to environmental change, and Arthur Georges (UCanberra) gave a presentation on using dartR for interfacing disgnosability with phylogenies to better inform species delimitation.

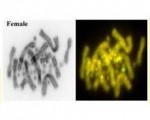
Day 3 focussed on landscape genetics, led by Bernd Gruber (UCanberra) and Renee Catullo (ANU), the use of resistance maps and simulation to address hypotheses on regional population structure. Floriaan Devloo-Delva (CSIRO) gave a talk on analysis options for identifying sex linked markers in sharks, and its implementation by Thierry in Radiator.

On Day 4, Brenna Forester (Colorado State) delivered a hands on session on outlier detection, data imputation, false-discovery rates, sourcing and screening environmental data, RDA (redundancy analysis), MEMs (Moran eigenvector maps), LFMM (latent factor mixed models) -- all contemporary approaches to landscape genetics, environmental associations and loci under selection.
The workshop finished up with topics on simulations for performance testing, parameter estimation, and developing null models delivered by Eric Archer (NOAA) using his R package strataG. An exciting step by step, hands-on exposition of simulation modelling. Mark Bravington (CSIRO) gave a presentation on "Close-kin genetics” as a way to understand animal ecology.