


















NewsBlog
eBook - Getting Started with dartRverse
Posted in Education and Outreach, Genomics onApr 08, 2026
New eBook comes on line -- Getting Started with dartRverse -- to help users begin working confidently with SNP data in the dartRverse environment.
Eureka Prize dartR
Posted in Education and Outreach onSep 07, 2025
We recently received the Eureka Prize for Excellence in Research Software for the dartR package. But what is dartR?
Lineages within Species or Species as Lineages
Posted in Turtle Research onMay 03, 2025
Taxonomy of Australian freshwater turtles is challenging. Here we present our latest findings, using SNPs and SilicoDArt molecular analysis, to examine species boundaries in the Emydura of northern Australia and Southern New Guinea.
Phylogeography of the Pignosed Turtle
Posted in Turtle Research onApr 07, 2025
In this paper we examine the phylogeographic genetic structure of the endangered pig-nosed turtle, the last remaining member of a once globally widespread family
Fossil feuds -- lavarackorum or oneiros?
Posted in Turtle Research onMar 17, 2023
New paper out in Vertebrate Zoology on the esoteric topic of whether the Australian Gulf Snapping Turtle is a living fossil (Elseya lavarackorum) or a living turtle with a recent fossil history (Elseya oneiros).
Dealing with Taxonomic Vandalism
Posted in Turtle Research onMar 28, 2022
The ASH List of Australian Species of Amphibian and Reptile was announced today. This is a major milestone that took several years to develop and achieve consensus. As one of the architects of the Positive List of journals (and other relevant works) that underpins the Species List, I outline here why I think it is a Positive List good idea. A Positive List distinguishes between science and non-science and may well put an end to the taxonomic instability that arises from the publication of poorly supported species descriptions promulgated outside the peer-reviewed scientific literature. Time will tell.
dartR Version 2 release
Posted in Genomics onMar 27, 2022
We are pleased to announce the release of dartR version 2. We have substantially increased the number of functions from 45 to 144 to enhance the user experience by extending plot customisation, function standardisation, increasing user support and function speed. dartR now provides many additional functions for importing data, exporting data and linking to other packages. Give it a try. And congratulations to the core development team Luis Mijangos, Bernd Gruber, Carlos Pacioni, Oliver Berry and Arthur Georges.
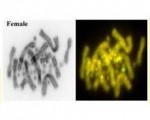
Sex in dragons -- reversal stable
Posted in Pogona Research onFeb 26, 2022
A combination of chromosomal sex determination and temperature-induced sex reversal appears to be evolutionarily stable based on evidence from an extensive field study of the central bearded dragon in central Queensland. Read more about it in a paper, led by Kris Wild, that came out today in Molecular Ecology.
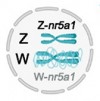
Sex in Dragons -- role for chromosomal conformation?
Posted in Pogona Research onJan 17, 2022
Reptiles have an extraordinary variety of mechanisms to determine sex. In an article that appeared in PNAS this week, we propose that altered configuration of the repeat-laden W chromosome affects the conformation of the primary transcript of a candidate sex determining gene to generate more diverse and potentially inhibitory W-borne isoforms that suppress testis determination. Epigenetics rules.
Epigenetics of the Dragon
Posted in Pogona Research onDec 15, 2021
Great to see that the antibodies for a range of sex related proteins developed for mammals work in our model species, the dragon lizard. Our article, led by Sarah Whiteley and as a collaboration between the QIMR Berghofer Institute for Medical Research, appeared today in the journal Biology of Reproduction.